Research Articles
Beyond the Mean: A Comprehensive Guide to Reaction Norm Analysis in Evolutionary Biology and Biomedical Research
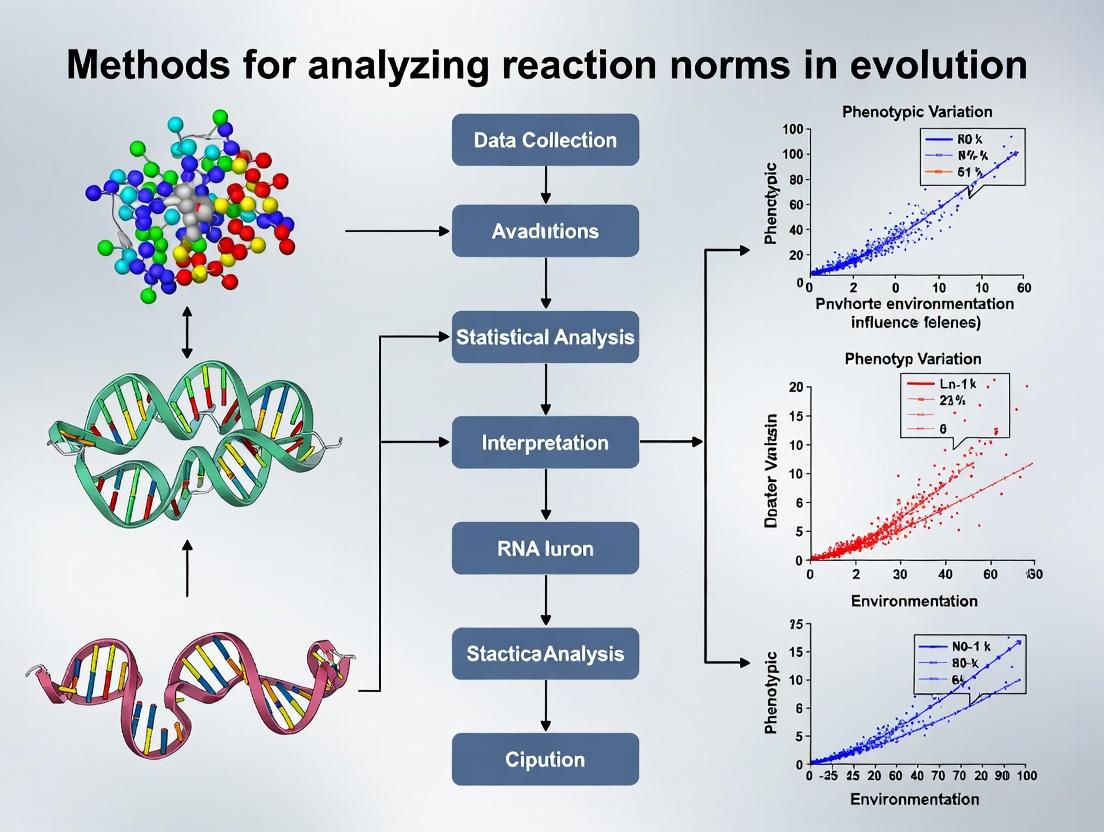
This article provides a detailed methodological guide for analyzing reaction norms—the patterns of phenotypic expression across environmental gradients—in evolutionary and biomedical contexts.
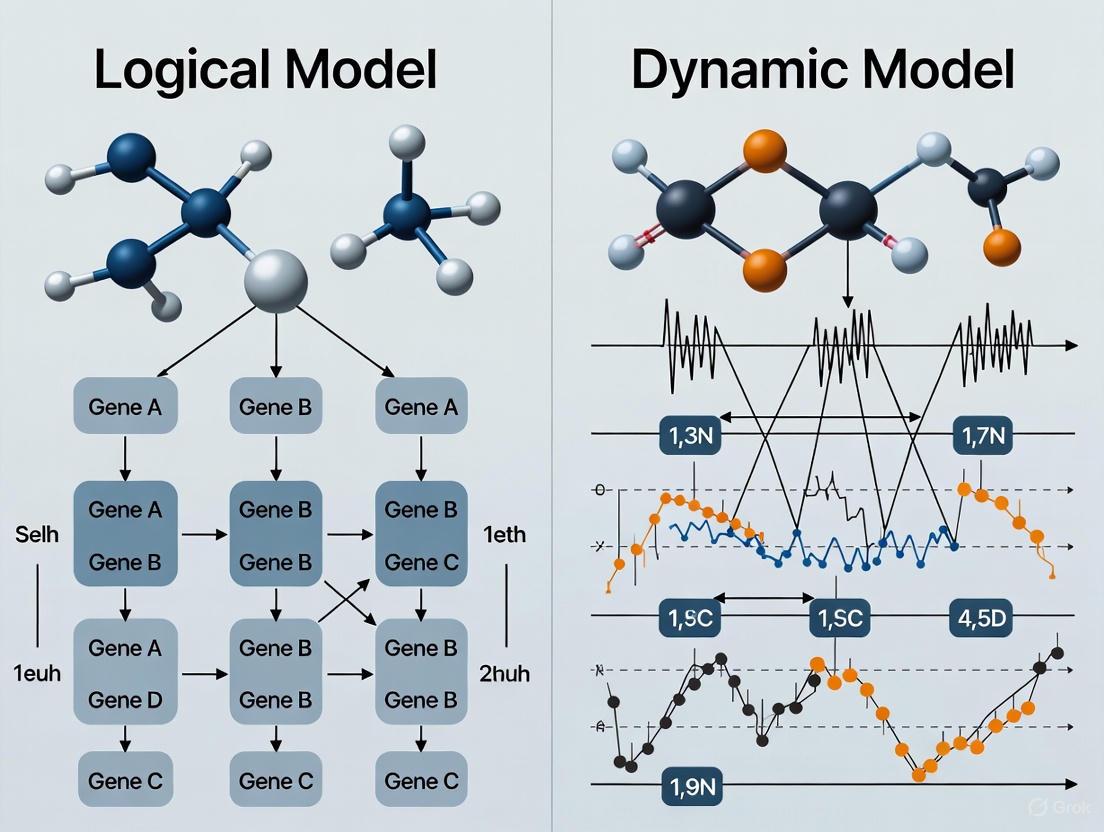
Logical vs Dynamic Models for Gene Networks: A Guide for Biomedical Researchers
This article provides a comprehensive comparison of logical and dynamic (quantitative) modeling frameworks for gene regulatory network (GRN) simulation, tailored for researchers, scientists, and drug development professionals.
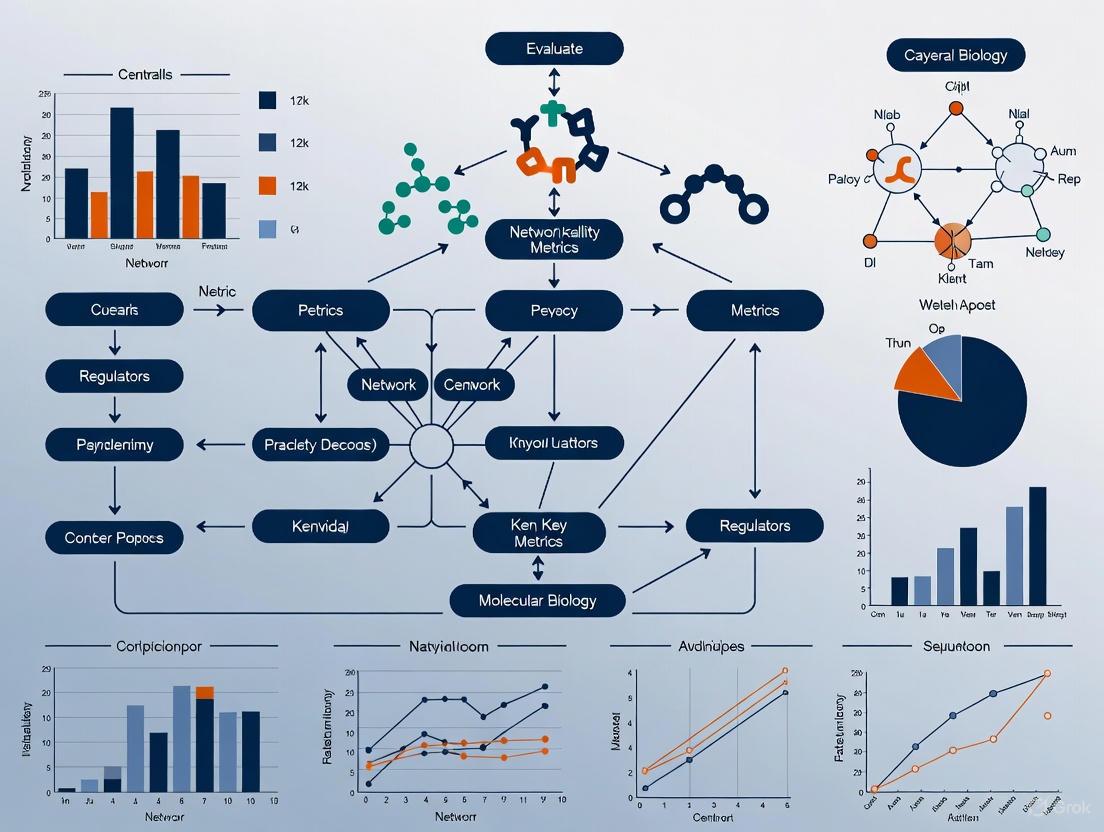
Network Centrality in Drug Discovery: A Guide to Identifying Key Regulatory Targets
This article provides a comprehensive guide for researchers and drug development professionals on the application of network centrality metrics to identify key regulatory targets in biological systems.
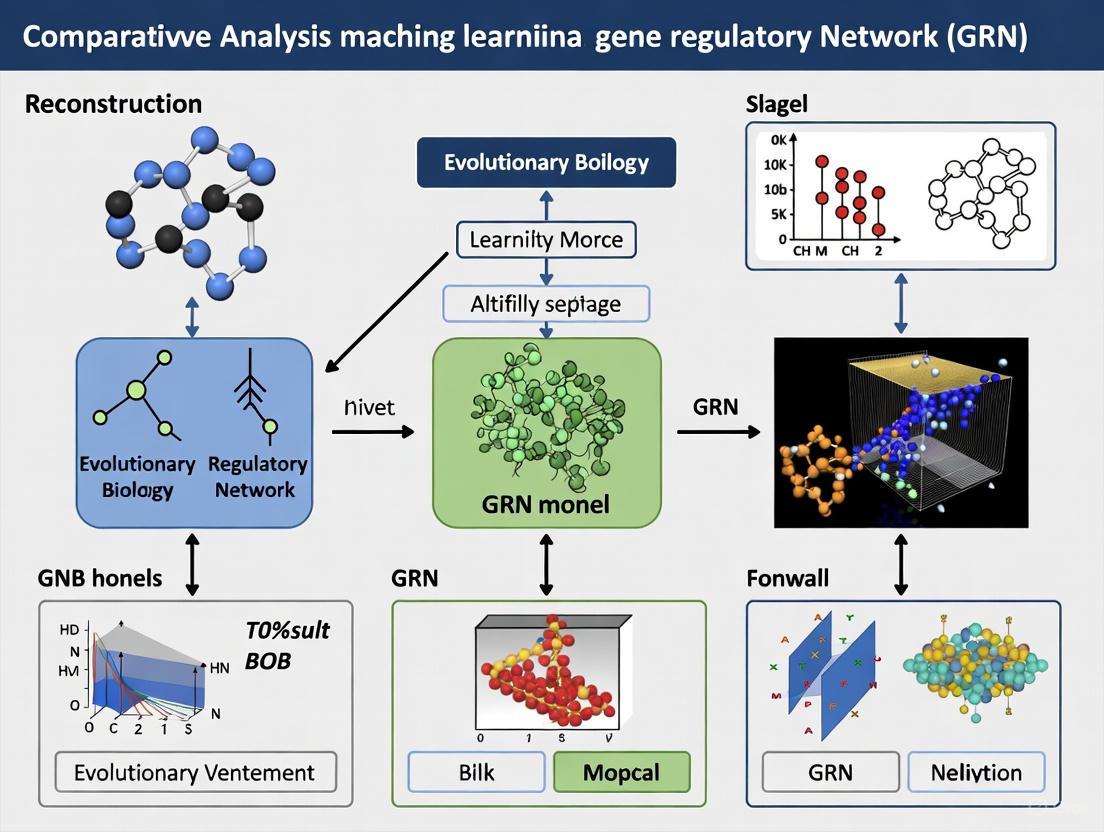
Machine Learning for Gene Regulatory Network Reconstruction: A Comparative Analysis of Methods, Challenges, and Future Directions
The reconstruction of Gene Regulatory Networks (GRNs) is fundamental for understanding cellular identity, disease mechanisms, and therapeutic target discovery.
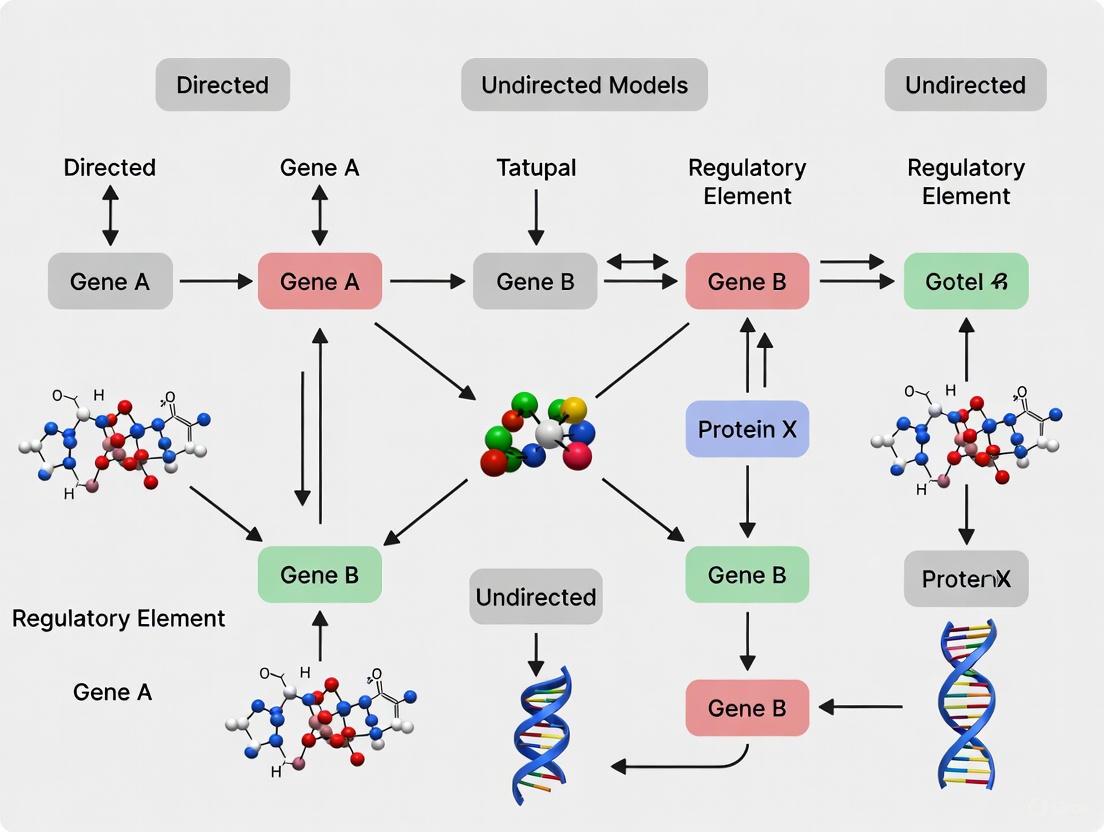
Directed vs. Undirected GRN Models: A Guide for Developmental Biology and Disease Research
This article provides a comprehensive comparison of directed and undirected graphical models for analyzing Gene Regulatory Networks (GRNs) in developmental processes.
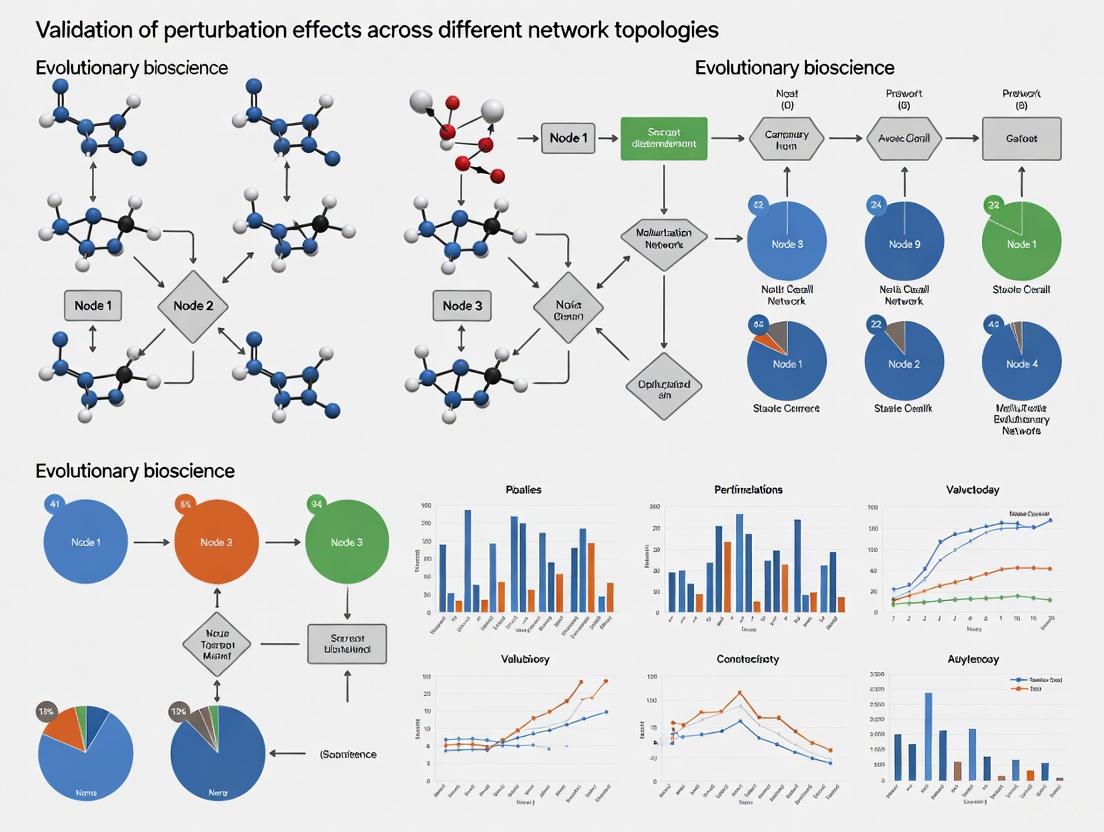
Validating Perturbation Effects Across Network Topologies: From Foundational Concepts to Clinical Applications
This article provides a comprehensive framework for researchers, scientists, and drug development professionals to validate perturbation effects across diverse network topologies.
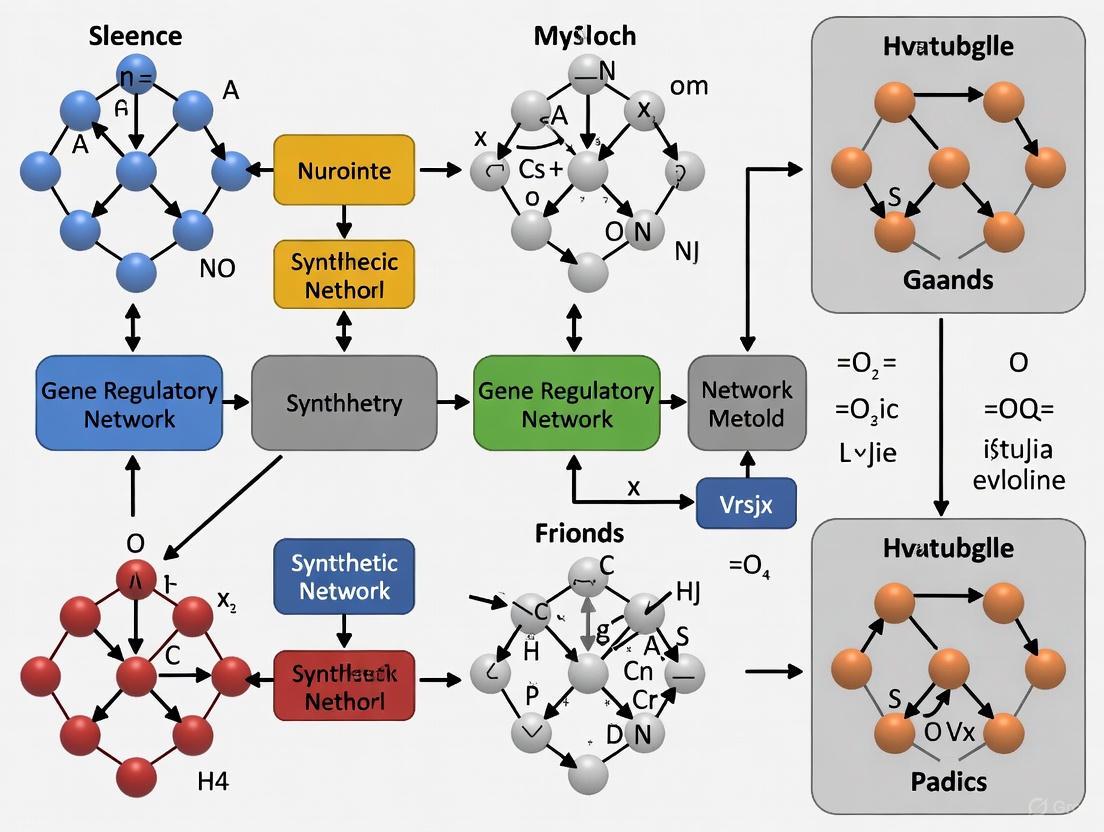
Benchmarking Gene Regulatory Network Inference: A Comprehensive Guide to Methods, Challenges, and Validation on Synthetic Networks
Inferring accurate Gene Regulatory Networks (GRNs) from high-throughput data is fundamental for understanding cellular mechanisms and advancing drug discovery.
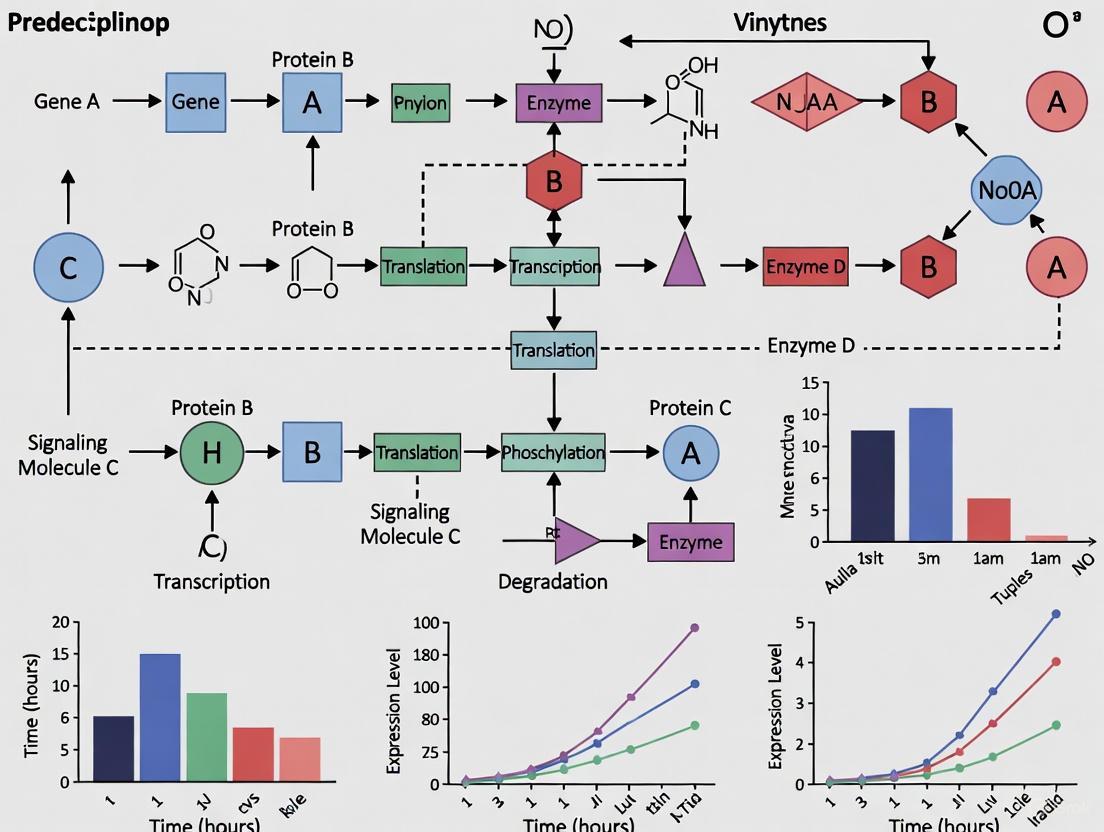
Key Challenges and Advanced Solutions in Predicting Bidirectional Regulation and Feedback Loops
This article explores the central challenges in modeling and predicting bidirectional regulation and feedback loops, dynamic systems fundamental to biology, from cellular decision-making to organism-level physiology.
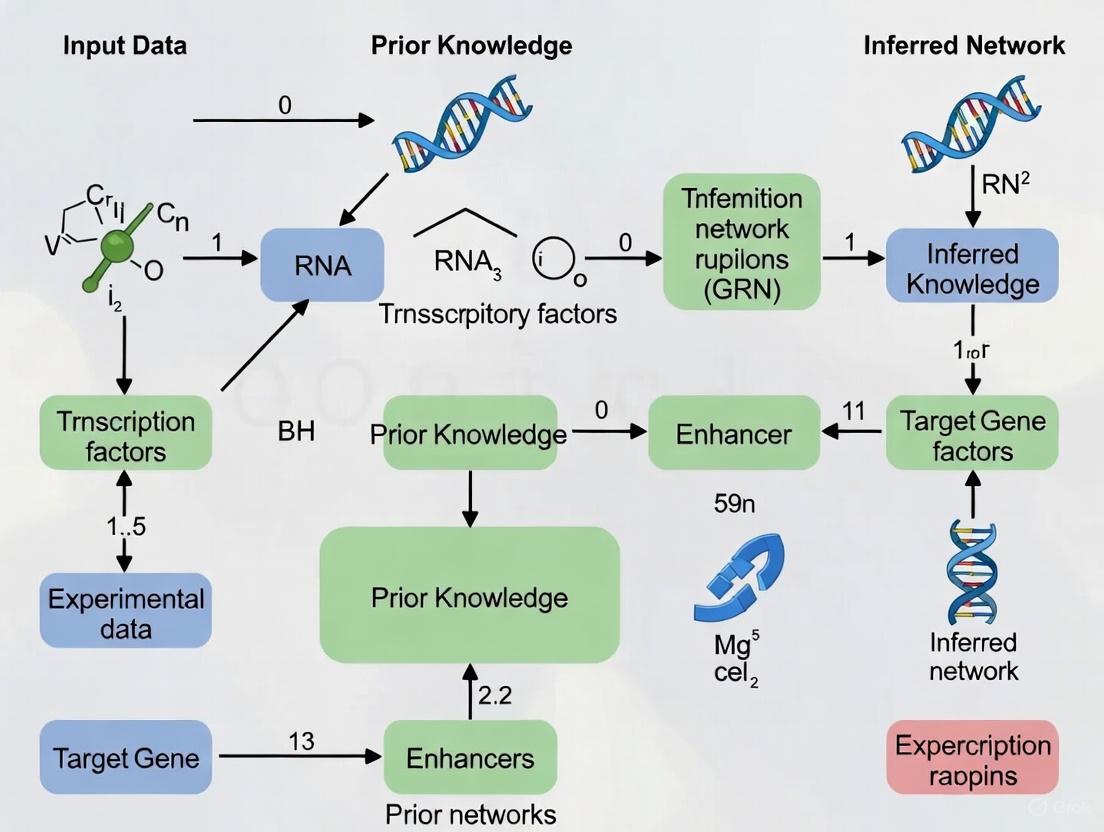
Integrating Prior Knowledge in GRN Inference: Advanced Strategies for Enhanced Accuracy and Biological Relevance
Accurately inferring Gene Regulatory Networks (GRNs) from single-cell RNA-sequencing data remains a significant challenge due to data sparsity and noise.
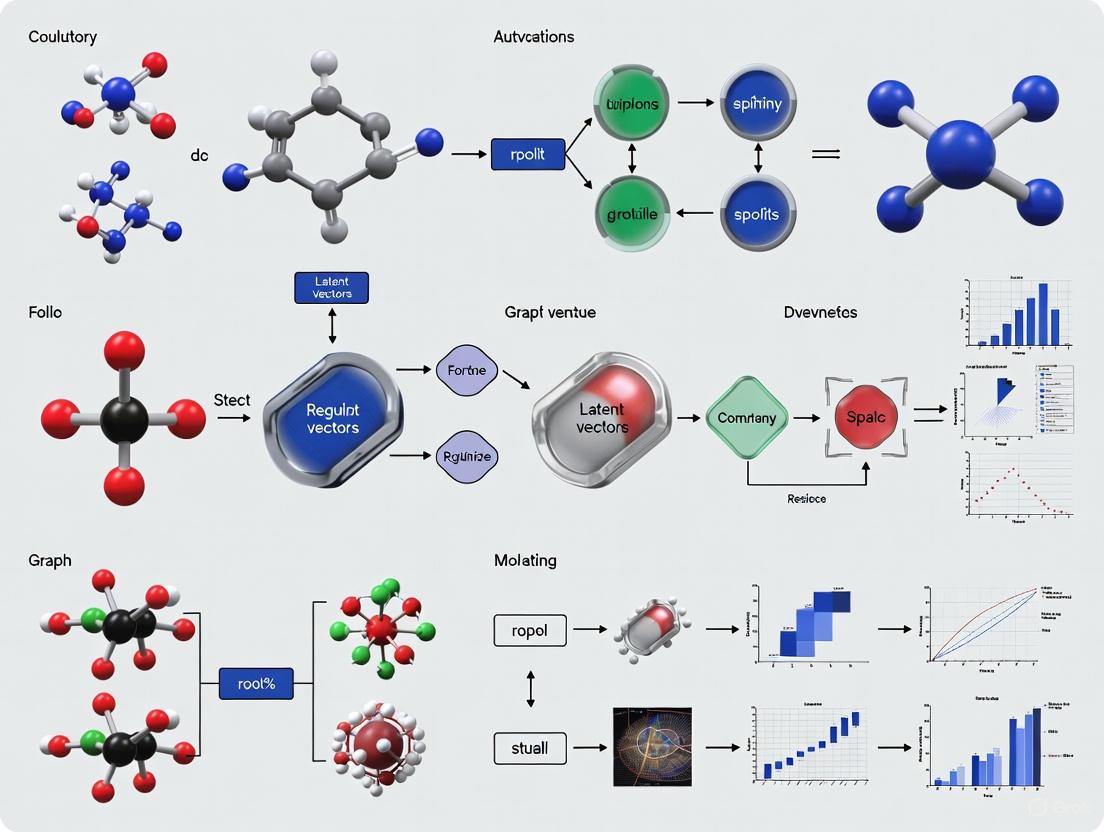
Advanced Latent Vector Regularization in Graph Autoencoders: Methods and Biomedical Applications
This article provides a comprehensive exploration of advanced regularization techniques for latent vectors in graph autoencoders, tailored for researchers and professionals in computational biology and drug discovery.